Static DICOMweb Files for Enhanced Performance
This section describes how to generate and serve static DICOMweb files, significantly improving the performance of your OHIF Viewer setup. These files are pre-processed and compressed, minimizing storage space and reducing serving time to the bare minimum (disk read and HTTP stream write).
Static-DICOMWeb Project
The core tool for this process is the static-wado project, available on GitHub:
This project contains two main components:
static-wado-creator: Converts raw DICOM files into a DICOMweb-compliant directory structure, optimizing them for efficient serving.static-wado-webserver: A simple web server specifically designed to serve the generated static DICOMweb files.
Prerequisites
- Node.js and npm (or yarn) installed on your system.
Installation
-
Clone the Repository:
git clone https://github.com/RadicalImaging/Static-DICOMWeb
cd Static-DICOMWeb -
Install Dependencies:
yarn install --frozen-lockfile
Generating Static DICOMweb Files
-
Prepare your DICOM data: Organize your DICOM files into a directory. For example
/Users/alireza/dicom/test-static-script/ACRIN-CT. -
Convert to DICOMweb Structure: Use the
mkdicomweb.jsscript from thestatic-wado-creatorpackage to create the DICOMweb directory:node packages/static-wado-creator/bin/mkdicomweb.js '/Users/alireza/dicom/test-static-script/ACRIN-CT' -o '/Users/alireza/dicom/test-static-script/output'- Replace:
/Users/alireza/dicom/test-static-script/ACRIN-CTwith the path to your directory of DICOM files./Users/alireza/dicom/test-static-script/outputwith your desired output directory for the DICOMweb structure.
This command will generate a directory structure similar to this in your output location:
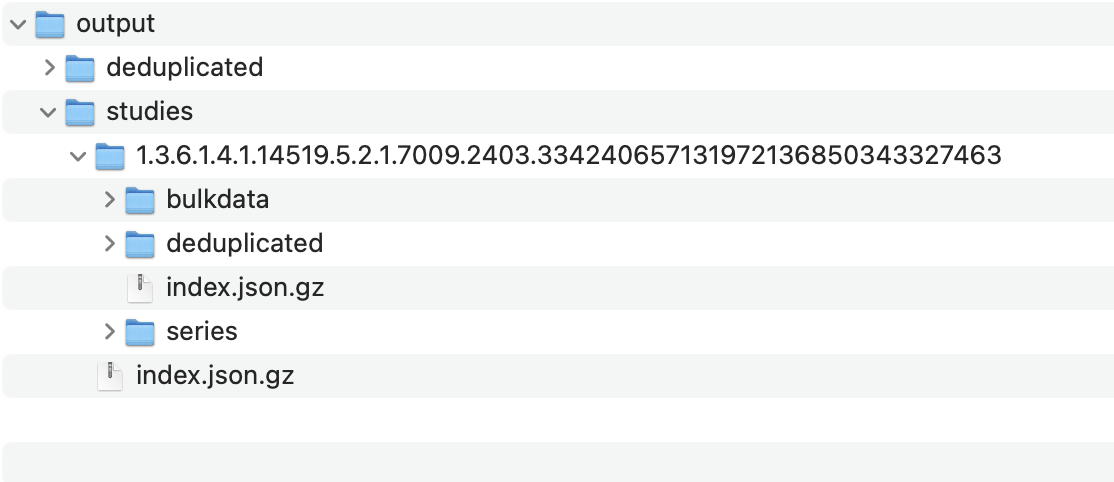
- Replace:
Serving Static Files with the Web Server
-
Start the Server:
Run the
dicomwebserver.mjsscript from thestatic-wado-webserverpackage, specifying the port and the DICOMweb directory:node packages/static-wado-webserver/bin/dicomwebserver.mjs -p 3001 -o /Users/alireza/dicom/test-static-script/output-p 3001: Sets the server to listen on port 3001. You can change this if needed.-o /Users/alireza/dicom/test-static-script/output: Specifies the path to your generated DICOMweb directory.
infoThe
-p(port) and-o(output directory) flags are used to configure the server.
Running OHIF Viewer with Static Data
-
Use the
local_static.jsConfiguration:Start the OHIF Viewer in development mode using the provided
local_static.jsconfiguration file:yarn dev:static -
Configuration Details:
The
local_static.jsconfiguration file is pre-configured to point to:qidoRoot: 'http://localhost:3001/dicomweb',
wadoRoot: 'http://localhost:3001/dicomweb',This matches the default port (3001) used by the
static-wado-webserver.infoIf you change the port or output directory when running the
static-wado-webserver, you must also update theqidoRootandwadoRootvalues in yourlocal_static.jsconfiguration file accordingly to ensure the OHIF Viewer can access the data.